NBO Scripts and Handy Applications

Welcome
... Natural bond orbital (NBO) analysis is a powerful method for formulating intuitive chemical hypotheses from otherwise complex quantum mechanical description of the molecular wavefunction. NBO description of chemical bonding conforms closely to the localized 2-electron/2-center concepts of Lewis and Pauling, both descriptions being a cornerstone of freshman chemistry ...
This website attempts to bring useful tools of Natural Bond Orbital analysis into one place while encouraging readers, researchers, and teachers to share their experience and ideas to further facilitate analysis, interpretation, and visualization of NBOs.
Over the past few years, I have been posting blog entries discussing approaches to visualization of Natural Bond Orbitals (NBO) in an open-source Java molecular viewer, Jmol. I wrote simple Java applications to assist with repetitive tasks accompanying the generation of Jmol scripts, macros and GENNBO inputs. Since blog space is not the best place for storing and tracking apps, it was only a matter of time before I decided to move the app files and links to static web site. Descriptions and examples still stay with the original blog posts.
To simplify the retrieval of NBO-related information from NBO result files, I opted for the Python language. Many scripts and Jupyter notebooks are available from the corresponding section of these pages. For more complex tasks involving multiple properties and visualizations, I prefer using the KNIME analytics and visualization environment. Several workflows are available for download from the KNIME section. To present results that include NBO orbital imagery, two methods employing jvxl and x3d files are described in building the "life" web-site display.
The main driver behind the Java apps, Jupyter Notebooks, and Java/Python workflows is to provide teachers, students, and researchers with simple tools for generating a wide variety of NBO results and visualizations without requirements of detailed knowledge of NBO keywords and Jmol syntax. It is mostly students of Organic Chemistry who will benefit from apps available at this web site, although I hope that professionals from other disciplines, such as computational chemistry and physical chemistry will find these topics useful as well.
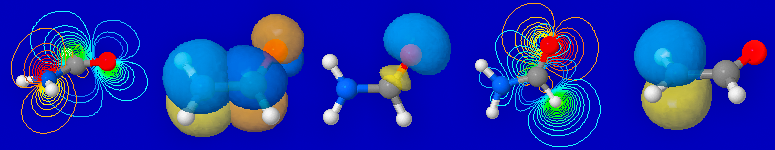
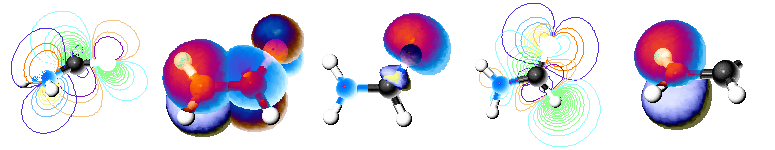
Jmol visualization of formamide
NBO Visualization Corner
Dynamic web visualization and jvxl surface files Latest
This blog explores Jmol-jvxl-based visualization of NBOs in a static website. Steps for getting the key elements for building the website are described in the Blog. The visualization is based on jvxl surface and structure files and it is supported by the JSmol. With the help of css, JavaScript, and JQuery, "gallery-type" interaction with molecular and surface models has been implemented.
The required jvxl files are generated using the Jmol-NBO Visualization Helper featured below. Jmol macro files allow loading and visualization of NBO archive files (.31-.46). A script is generated that writes the NBO surfaces into the corresponding jvxl files. Those are then loaded into the web interface with the assistance of JSmol serving as a client backend.
All files described in the post are available from the Download section of the Blog. The site was recently re-build using the Bootstrap framework and LESS dynamic style sheet language. It is fully responsive and supports devices, such as phones and tablets.
The accompanying demo webpage can be viewed
here.
Web Visualization of NBOs in x3d Format Latest
In this post, visualization capabilities of Jmol are exploited for generating the orbital imagery that uses open-source x3d format. X3D standard is XML-based and supports visualization of 3D computer graphics.
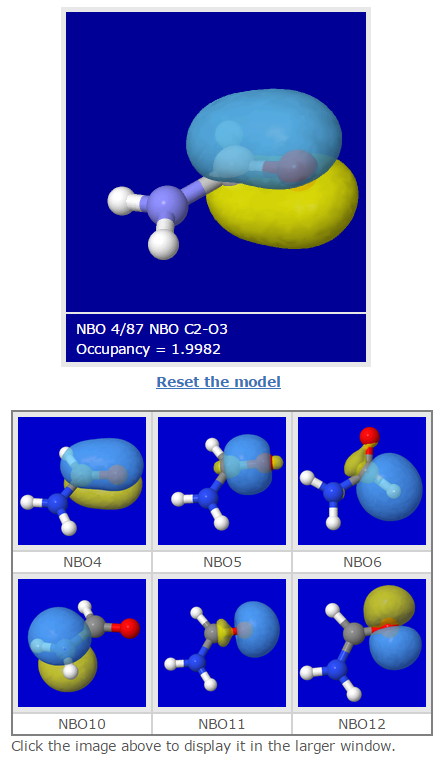
As in the jvxl-related Blog above, Jmol-NBO Visualization Helper was used to create Jmol macro files and scripts that assist in writing the .x3d files. These are used to load 3D images in the webpage through the similar "gallery-type" interface.
The accompanying demo webpage can be viewed here.
... discussed is embedding NBO imagery into a Web page using the Jmol-generated x3d and jvxl file formats. While the web implementation ofx3dfiles does not require the Jmol app or JavaScript support, successful implementation ofjvxlfiles takes advantage of both. Web page template and required resources can be downloaded and adapted to a variety of presentation settings.
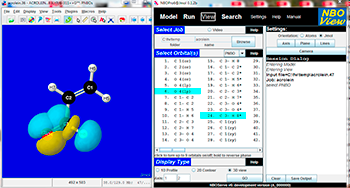
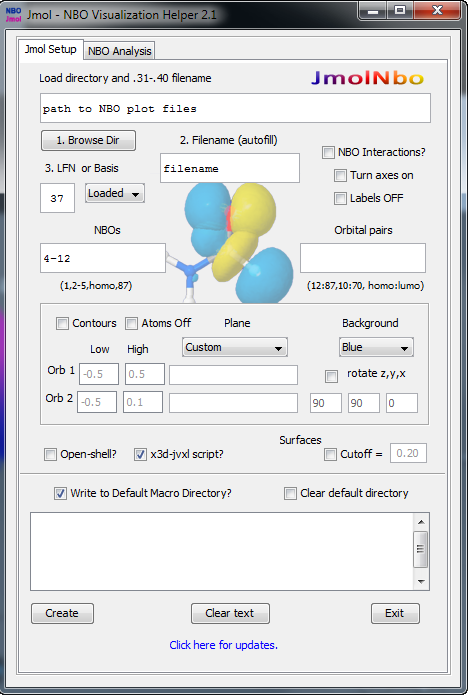
Visualization of Natural Bond Orbitals is best accomplished in the Jmol Viewer. JmolNBO Visualization Helper (image of the GUI on the right) assists with setting the necessary parameters to display AO->->NLMO orbitals. Other methods of visualization can be found on the Web, such as GaussView and Molden or Molekel.
NBO Highlights
Featured review article
An excellent review explaining practical details of NBO analysis. Published in: WIREs Comput Mol Sci 2012, 2: 1–42
NBO7 Announced
Following years of program development and cooperation with affiliated ESS providers, leading members of the NBO Team (Eric Glendening, Frank Weinhold, Clark Landis) unveiled the new nbo7.chem.wisc.edu website and announced public availability of the NBO 7.0 program, as well as enhanced NBOPro@Jmol program.